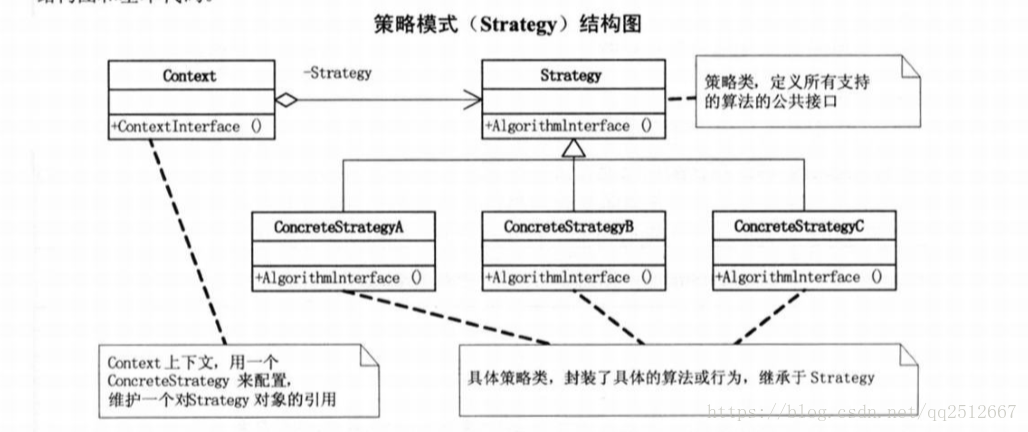
代码参考自:https://github.com/orvindemsy/MI-BCI-CSP, 做了整理与封装,更方便使用
输入数据格式为:x_shape = [trial, channal, timepoint], y_shape = [trial]
python">from mimetypes import init
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import scipy.io as io
from pandas import DataFrame as df
from scipy.linalg import inv, sqrtm
from sklearn.metrics import accuracy_score, classification_report
from sklearn.model_selection import (GridSearchCV, KFold, cross_val_score,
train_test_split)
from sklearn.svm import SVC
def compute_cov(EEG_data):
'''
INPUT:
EEG_data : EEG_data in shape T x N x S
OUTPUT:
avg_cov : covariance matrix of averaged over all trials
'''
cov = []
for i in range(EEG_data.shape[0]):
cov.append(EEG_data[i]@EEG_data[i].T/np.trace(EEG_data[i]@EEG_data[i].T))
cov = np.mean(np.array(cov), 0)
return cov
def decompose_cov(avg_cov):
'''
This function will decompose average covariance matrix of one class of each subject into
eigenvalues denoted by lambda and eigenvector denoted by V
Both will be in descending order
Parameter:
avgCov = the averaged covariance of one class
Return:
λ_dsc and V_dsc, i.e. eigenvalues and eigenvector in descending order
'''
λ, V = np.linalg.eig(avg_cov)
λ_dsc = np.sort(λ)[::-1] # Sort eigenvalue descending order, default is ascending order sort
idx_dsc = np.argsort(λ)[::-1] # Find index in descending order
V_dsc = V[:, idx_dsc] # Sort eigenvectors descending order
λ_dsc = np.diag(λ_dsc) # Diagonalize λ_dsc
return λ_dsc, V_dsc
def white_matrix(λ_dsc, V_dsc):
'''
'''
λ_dsc_sqr = sqrtm(inv(λ_dsc))
P = (λ_dsc_sqr)@(V_dsc.T)
return P
def compute_S(avg_Cov, white):
'''
This function will compute S matrix, S = P * C * P.T
INPUT:
avg_Cov: averaged covariance of one class, dimension N x N, where N is number of electrodes
white: the whitening transformation matrix
OUTPUT:
S
'''
S = white@avg_Cov@white.T
return S
def decompose_S(S_one_class, order='descending'):
'''
This function will decompose the S matrix of one class to get the eigen vector
Both eigenvector will be the same but in opposite order
i.e the highest eigenvector in S left will be equal to lowest eigenvector in S right matrix
'''
# Decompose S
λ, B = np.linalg.eig(S_one_class)
# Sort eigenvalues either descending or ascending
if order == 'ascending':
idx = λ.argsort() # Use this index to sort eigenvector smallest -> largest
elif order == 'descending':
idx = λ.argsort()[::-1] # Use this index to sort eigenvector largest -> smallest
else:
print('Wrong order input')
λ = λ[idx]
B = B[:, idx]
return B, λ
def spatial_filter(B, P):
'''
Will compute projection matrix using the following equation:
W = B' @ P
INPUT:
B: the eigenvector either left or right class, choose one, size N x N, N is number of electrodes
P: white matrix in size of N x N
OUTPUT:
W spatial filter to filter EEG
'''
return (B.T@P)
def compute_Z(W, E, m):
'''
Will compute the Z
Z = W @ E,
E is in the shape of N x M, N is number of electrodes, M is sample
In application, E has nth trial, so there will be n numbers of Z
Z, in each trial will have dimension of m x M,
where m is the first and last m rows of W, corresponds to smallest and largest eigenvalues
'''
Z = []
W = np.delete(W, np.s_[m:-m:], 0)
for i in range(E.shape[0]):
Z.append(W @ E[i])
return np.array(Z)
def feat_vector(Z):
'''
Will compute the feature vector of Z matrix
INPUT:
Z : projected EEG shape of T x N x S
OUTPUT:
feat : feature vector shape of T x m
T = trial
N = channel
S = sample
m = number of filter
'''
feat = []
for i in range(Z.shape[0]):
var = np.var(Z[i], ddof=1, axis=1)
varsum = np.sum(var)
feat.append(np.log10(var/varsum))
return np.array(feat)
class CSP():
def __init__(self,m = 2):
super(CSP,self).__init__()
self.m = m
self. W = None
def fit(self,x_train,y_train):
# 分成左右手的数据
x_train_l = x_train[y_train==0]
x_train_r = x_train[y_train==1]
# 算平均协方差阵
C_l = compute_cov(x_train_l)
C_r = compute_cov(x_train_r)
C_c = C_l+C_r
# 白化
eigval, eigvec = decompose_cov(C_c)
P = white_matrix(eigval,eigvec)
# 计算S_l,S_r
S_l = compute_S(C_l, P)
S_r = compute_S(C_r, P)
# 分解S_l,S_r
S_l_eigvec, S_l_eigval = decompose_S(S_l, 'descending')
S_r_eigvec, S_r_eigval = decompose_S(S_r, 'ascending')
# 计算W
self.W = spatial_filter(S_l_eigvec,P)
def transform(self,x_train):
Z_train = compute_Z(self.W,x_train,self.m)
feat_train = feat_vector(Z_train)
return feat_train
def fit_transform(self,x_train,y_trian):
self.fit(x_train,y_trian)
feat_train = self.transform(x_train)
return feat_train